scTour: a deep learning architecture for robust inference and accurate prediction of cellular dynamics
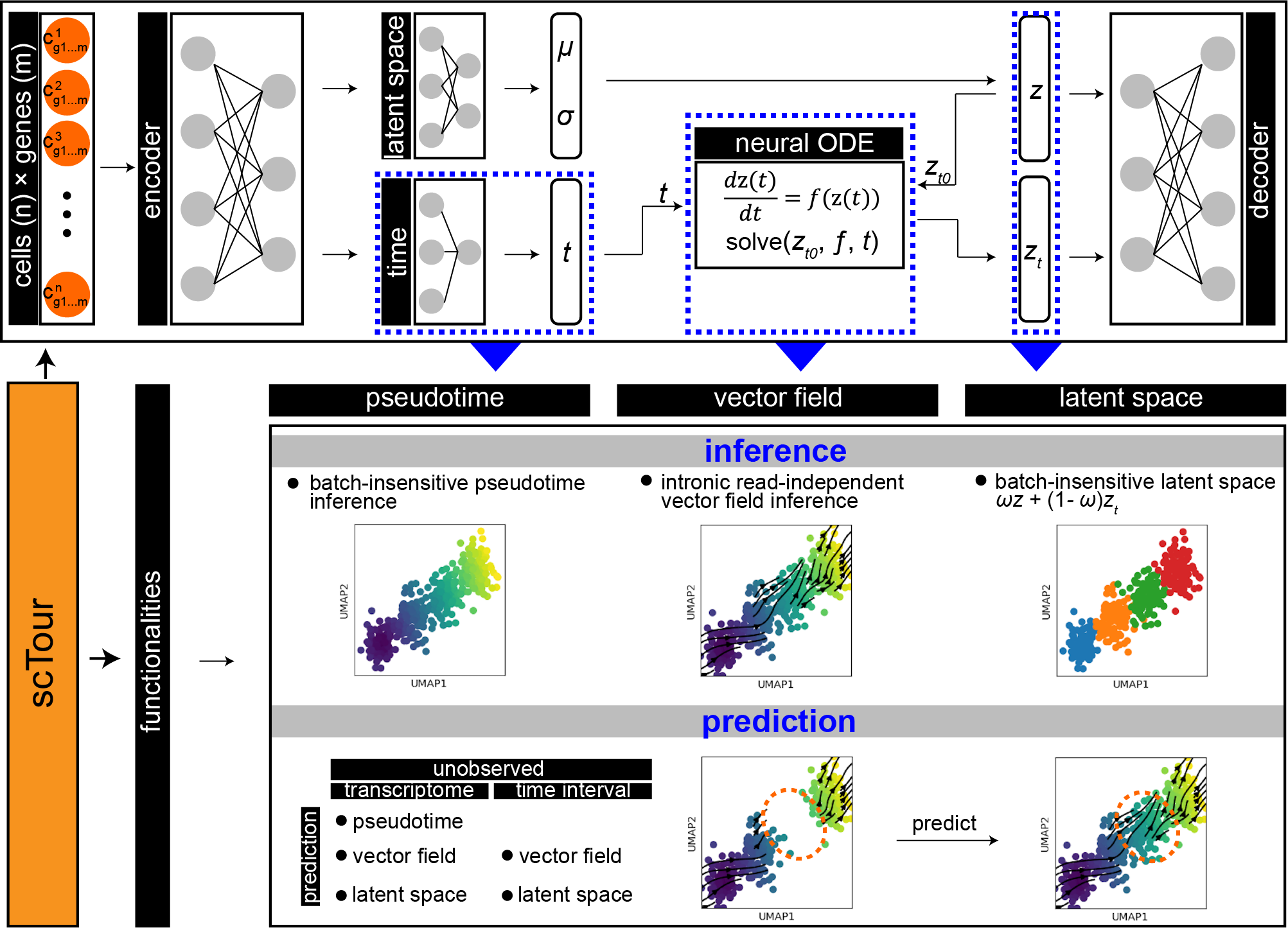
scTour is an innovative and comprehensive method for dissecting cellular dynamics by analysing datasets derived from single-cell genomics.
It provides a unifying framework to depict the full picture of developmental processes from multiple angles including the developmental pseudotime, vector field and latent space.
It further generalises these functionalities to a multi-task architecture for within-dataset inference and cross-dataset prediction of cellular dynamics in a batch-insensitive manner.
Key features
cell pseudotime estimation with no need for specifying starting cells.
transcriptomic vector field inference with no discrimination between spliced and unspliced mRNAs.
latent space mapping by combining intrinsic transcriptomic structure with extrinsic pseudotime ordering.
model-based prediction of pseudotime, vector field, and latent space for query cells/datasets/time intervals.
insensitive to batch effects; robust to cell subsampling; scalable to large datasets.
Installation
scTour requires Python ≥ 3.7:
pip install sctour
Reference
Qian Li, scTour: a deep learning architecture for robust inference and accurate prediction of cellular dynamics, 2023, Genome Biology.
